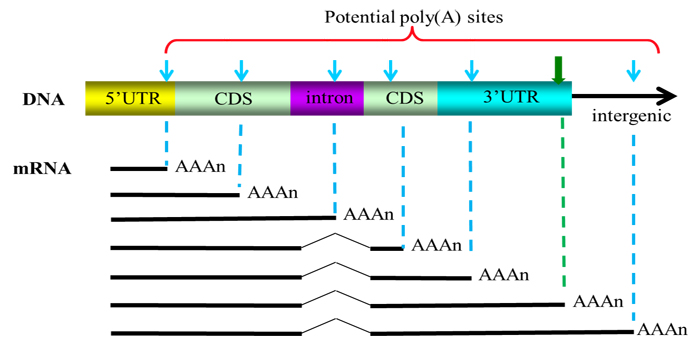
Intronic poly(A) sites in human genes. (A) Schematic of poly(A) sites... | Download Scientific Diagram
PLOS ONE: Reprogramming of 3′ Untranslated Regions of mRNAs by Alternative Polyadenylation in Generation of Pluripotent Stem Cells from Different Cell Types

Figure 1-1 from Deconvoluting Cell-Type Specific 3'UTR Isoform Expression in the Adult and Developing Cerebellum | Semantic Scholar
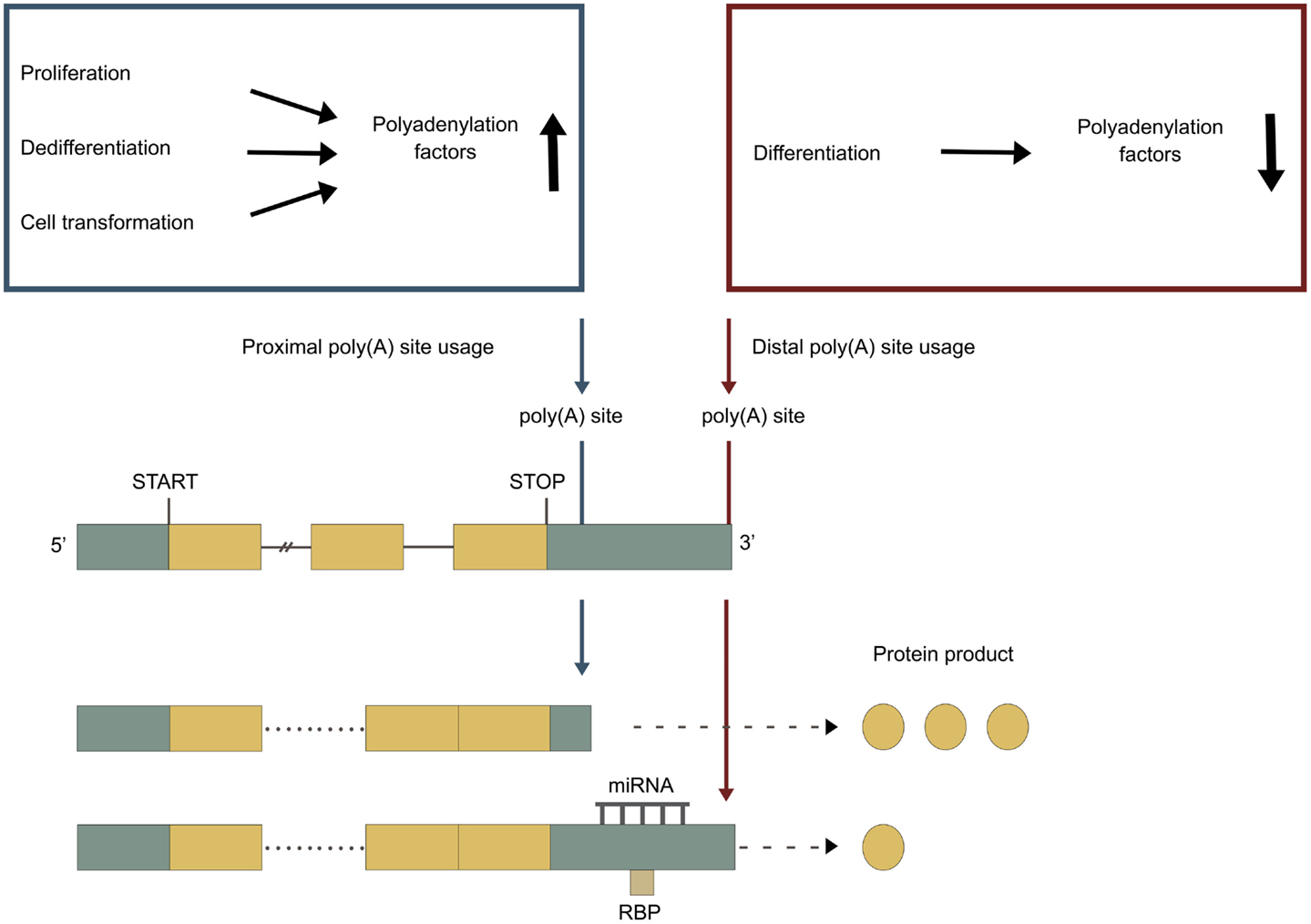
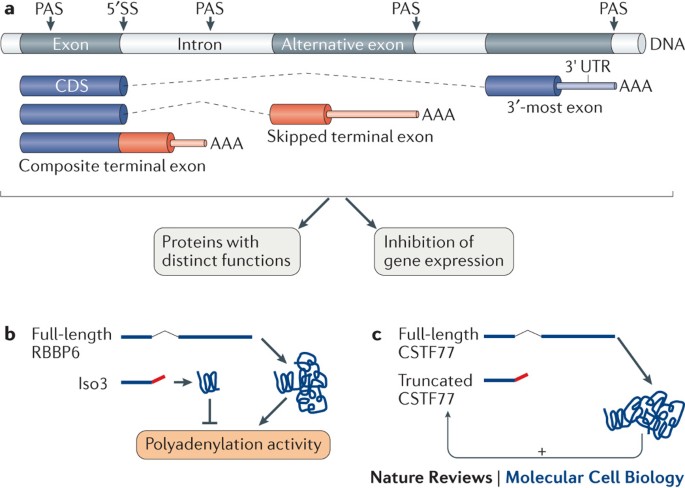
Frontiers | Alterations in Polyadenylation and Its Implications for Endocrine Disease | Endocrinology
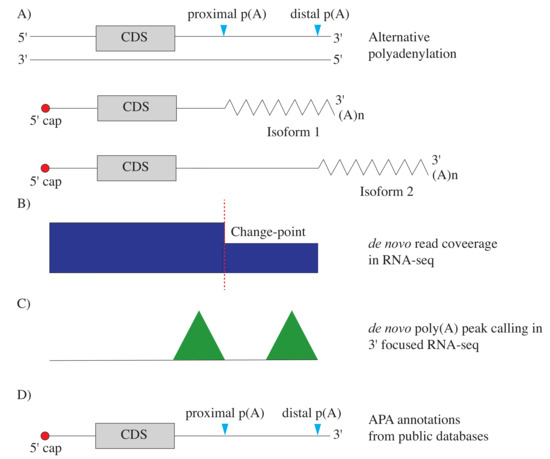
IJMS | Free Full-Text | The Detection and Bioinformatic Analysis of Alternative 3′ UTR Isoforms as Potential Cancer Biomarkers | HTML

Poly(A)-ClickSeq: click-chemistry for next-generation 3´-end sequencing without RNA enrichment or fragmentation | bioRxiv

Genome-wide landscape of polyadenylation in Arabidopsis provides evidence for extensive alternative polyadenylation | PNAS

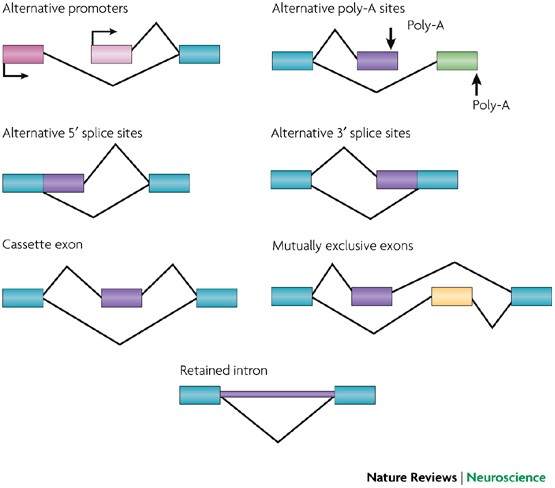
Four types of alternative polyadenylation events. (A) Tandem 3 0 UTR... | Download Scientific Diagram

Figure 9 from Formation of mRNA 3′ Ends in Eukaryotes: Mechanism, Regulation, and Interrelationships with Other Steps in mRNA Synthesis | Semantic Scholar

Progressive lengthening of 3′ untranslated regions of mRNAs by alternative polyadenylation during mouse embryonic development | PNAS

The Poly(A)-Binding Protein Nuclear 1 Suppresses Alternative Cleavage and Polyadenylation Sites: Cell